Retracted!
Do not use this motif!
Do not use this motif!
Model info
| Transcription factor | Evx2 | ||||||||
| Model | EVX2_MOUSE.H11DI.0.A RETRACTED!!! | ||||||||
| Model type | Dinucleotide PWM | ||||||||
| LOGO | 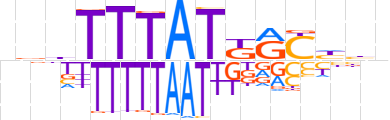 | ||||||||
| LOGO (reverse complement) |  | ||||||||
| Data source | ChIP-Seq | ||||||||
| Model release | HOCOMOCOv11 | ||||||||
| Model length | 14 | ||||||||
| Quality | A | ||||||||
| Motif rank | 0 | ||||||||
| Consensus | nbbTTTATKGYhbb | ||||||||
| Best auROC (human) | 0.934 | ||||||||
| Best auROC (mouse) | 0.971 | ||||||||
| Peak sets in benchmark (human) | 11 | ||||||||
| Peak sets in benchmark (mouse) | 4 | ||||||||
| Aligned words | 250 | ||||||||
| TF family | HOX-related factors {3.1.1} | ||||||||
| TF subfamily | EVX (Even-skipped homolog) {3.1.1.10} | ||||||||
| MGI | MGI:95462 | ||||||||
| EntrezGene | GeneID:14029 (SSTAR profile) | ||||||||
| UniProt ID | EVX2_MOUSE | ||||||||
| UniProt AC | P49749 (TFClass) | ||||||||
| Comment | Retracted motif! | ||||||||
| Downloads | pcm
pwm alignment threshold to P-value grid | ||||||||
| Standard thresholds |
| ||||||||
| GTEx tissue expression atlas | Evx2 expression | ||||||||
| Motifs in JASPAR |
PCM
| AA | AC | AG | AT | CA | CC | CG | CT | GA | GC | GG | GT | TA | TC | TG | TT | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 01 | 8.0 | 12.0 | 35.0 | 21.0 | 7.0 | 8.0 | 9.0 | 16.0 | 8.0 | 13.0 | 25.0 | 20.0 | 10.0 | 14.0 | 25.0 | 8.0 |
| 02 | 5.0 | 17.0 | 3.0 | 8.0 | 12.0 | 6.0 | 8.0 | 21.0 | 15.0 | 12.0 | 16.0 | 51.0 | 6.0 | 10.0 | 21.0 | 28.0 |
| 03 | 0.0 | 0.0 | 1.0 | 37.0 | 0.0 | 3.0 | 0.0 | 42.0 | 0.0 | 0.0 | 1.0 | 47.0 | 0.0 | 1.0 | 5.0 | 102.0 |
| 04 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 | 3.0 | 0.0 | 0.0 | 0.0 | 7.0 | 4.0 | 1.0 | 0.0 | 223.0 |
| 05 | 0.0 | 0.0 | 1.0 | 3.0 | 1.0 | 0.0 | 0.0 | 1.0 | 0.0 | 0.0 | 0.0 | 0.0 | 3.0 | 0.0 | 10.0 | 220.0 |
| 06 | 4.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 11.0 | 0.0 | 0.0 | 0.0 | 224.0 | 0.0 | 0.0 | 0.0 |
| 07 | 0.0 | 10.0 | 0.0 | 229.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 |
| 08 | 0.0 | 0.0 | 0.0 | 0.0 | 3.0 | 0.0 | 5.0 | 2.0 | 0.0 | 0.0 | 0.0 | 0.0 | 6.0 | 5.0 | 118.0 | 100.0 |
| 09 | 2.0 | 0.0 | 7.0 | 0.0 | 5.0 | 0.0 | 0.0 | 0.0 | 66.0 | 0.0 | 57.0 | 0.0 | 26.0 | 0.0 | 76.0 | 0.0 |
| 10 | 0.0 | 77.0 | 10.0 | 12.0 | 0.0 | 0.0 | 0.0 | 0.0 | 5.0 | 110.0 | 12.0 | 13.0 | 0.0 | 0.0 | 0.0 | 0.0 |
| 11 | 0.0 | 2.0 | 1.0 | 2.0 | 29.0 | 83.0 | 3.0 | 72.0 | 1.0 | 9.0 | 3.0 | 9.0 | 4.0 | 7.0 | 5.0 | 9.0 |
| 12 | 0.0 | 12.0 | 18.0 | 4.0 | 14.0 | 49.0 | 3.0 | 35.0 | 1.0 | 7.0 | 1.0 | 3.0 | 8.0 | 21.0 | 32.0 | 31.0 |
| 13 | 1.0 | 7.0 | 4.0 | 11.0 | 20.0 | 26.0 | 5.0 | 38.0 | 3.0 | 15.0 | 18.0 | 18.0 | 15.0 | 16.0 | 16.0 | 26.0 |
PWM
| AA | AC | AG | AT | CA | CC | CG | CT | GA | GC | GG | GT | TA | TC | TG | TT | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 01 | -0.605 | -0.213 | 0.839 | 0.334 | -0.733 | -0.605 | -0.492 | 0.067 | -0.605 | -0.136 | 0.506 | 0.286 | -0.39 | -0.063 | 0.506 | -0.605 |
| 02 | -1.051 | 0.127 | -1.52 | -0.605 | -0.213 | -0.879 | -0.605 | 0.334 | 0.004 | -0.213 | 0.067 | 1.212 | -0.879 | -0.39 | 0.334 | 0.618 |
| 03 | -3.799 | -3.799 | -2.432 | 0.894 | -3.799 | -1.52 | -3.799 | 1.019 | -3.799 | -3.799 | -2.432 | 1.131 | -3.799 | -2.432 | -1.051 | 1.902 |
| 04 | -3.799 | -3.799 | -3.799 | -3.799 | -3.799 | -2.432 | -3.799 | -1.52 | -3.799 | -3.799 | -3.799 | -0.733 | -1.258 | -2.432 | -3.799 | 2.682 |
| 05 | -3.799 | -3.799 | -2.432 | -1.52 | -2.432 | -3.799 | -3.799 | -2.432 | -3.799 | -3.799 | -3.799 | -3.799 | -1.52 | -3.799 | -0.39 | 2.669 |
| 06 | -1.258 | -3.799 | -3.799 | -3.799 | -3.799 | -3.799 | -3.799 | -3.799 | -0.298 | -3.799 | -3.799 | -3.799 | 2.687 | -3.799 | -3.799 | -3.799 |
| 07 | -3.799 | -0.39 | -3.799 | 2.709 | -3.799 | -3.799 | -3.799 | -3.799 | -3.799 | -3.799 | -3.799 | -3.799 | -3.799 | -3.799 | -3.799 | -3.799 |
| 08 | -3.799 | -3.799 | -3.799 | -3.799 | -1.52 | -3.799 | -1.051 | -1.875 | -3.799 | -3.799 | -3.799 | -3.799 | -0.879 | -1.051 | 2.047 | 1.882 |
| 09 | -1.875 | -3.799 | -0.733 | -3.799 | -1.051 | -3.799 | -3.799 | -3.799 | 1.468 | -3.799 | 1.323 | -3.799 | 0.545 | -3.799 | 1.609 | -3.799 |
| 10 | -3.799 | 1.622 | -0.39 | -0.213 | -3.799 | -3.799 | -3.799 | -3.799 | -1.051 | 1.977 | -0.213 | -0.136 | -3.799 | -3.799 | -3.799 | -3.799 |
| 11 | -3.799 | -1.875 | -2.432 | -1.875 | 0.652 | 1.696 | -1.52 | 1.555 | -2.432 | -0.492 | -1.52 | -0.492 | -1.258 | -0.733 | -1.051 | -0.492 |
| 12 | -3.799 | -0.213 | 0.183 | -1.258 | -0.063 | 1.172 | -1.52 | 0.839 | -2.432 | -0.733 | -2.432 | -1.52 | -0.605 | 0.334 | 0.75 | 0.718 |
| 13 | -2.432 | -0.733 | -1.258 | -0.298 | 0.286 | 0.545 | -1.051 | 0.92 | -1.52 | 0.004 | 0.183 | 0.183 | 0.004 | 0.067 | 0.067 | 0.545 |