Retracted!
Do not use this motif!
Do not use this motif!
Model info
| Transcription factor | NFKB1 (GeneCards) | ||||||||
| Model | NFKB1_HUMAN.H11DI.0.A RETRACTED!!! | ||||||||
| Model type | Dinucleotide PWM | ||||||||
| LOGO |  | ||||||||
| LOGO (reverse complement) | 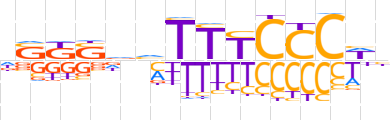 | ||||||||
| Data source | ChIP-Seq | ||||||||
| Model release | HOCOMOCOv11 | ||||||||
| Model length | 14 | ||||||||
| Quality | A | ||||||||
| Motif rank | 0 | ||||||||
| Consensus | nWGGGRAAbhSMSh | ||||||||
| Best auROC (human) | 0.867 | ||||||||
| Best auROC (mouse) | 0.958 | ||||||||
| Peak sets in benchmark (human) | 7 | ||||||||
| Peak sets in benchmark (mouse) | 16 | ||||||||
| Aligned words | 319 | ||||||||
| TF family | NF-kappaB-related factors {6.1.1} | ||||||||
| TF subfamily | NF-kappaB p50 subunit-like factors {6.1.1.1} | ||||||||
| HGNC | HGNC:7794 | ||||||||
| EntrezGene | GeneID:4790 (SSTAR profile) | ||||||||
| UniProt ID | NFKB1_HUMAN | ||||||||
| UniProt AC | P19838 (TFClass) | ||||||||
| Comment | Retracted motif! | ||||||||
| Downloads | pcm
pwm alignment threshold to P-value grid | ||||||||
| Standard thresholds |
| ||||||||
| GTEx tissue expression atlas | NFKB1 expression | ||||||||
| ReMap ChIP-seq dataset list | NFKB1 datasets | ||||||||
| Motifs in JASPAR |
PCM
| AA | AC | AG | AT | CA | CC | CG | CT | GA | GC | GG | GT | TA | TC | TG | TT | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 01 | 47.0 | 4.0 | 6.0 | 18.0 | 25.0 | 4.0 | 9.0 | 18.0 | 51.0 | 2.0 | 1.0 | 15.0 | 26.0 | 1.0 | 22.0 | 14.0 |
| 02 | 2.0 | 3.0 | 139.0 | 5.0 | 0.0 | 0.0 | 11.0 | 0.0 | 0.0 | 1.0 | 36.0 | 1.0 | 1.0 | 1.0 | 61.0 | 2.0 |
| 03 | 0.0 | 0.0 | 3.0 | 0.0 | 1.0 | 0.0 | 3.0 | 1.0 | 46.0 | 1.0 | 198.0 | 2.0 | 2.0 | 0.0 | 6.0 | 0.0 |
| 04 | 4.0 | 0.0 | 45.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.0 | 20.0 | 0.0 | 188.0 | 2.0 | 0.0 | 0.0 | 2.0 | 1.0 |
| 05 | 17.0 | 0.0 | 6.0 | 1.0 | 0.0 | 0.0 | 0.0 | 0.0 | 171.0 | 3.0 | 57.0 | 5.0 | 3.0 | 0.0 | 0.0 | 0.0 |
| 06 | 156.0 | 2.0 | 27.0 | 6.0 | 1.0 | 0.0 | 2.0 | 0.0 | 62.0 | 0.0 | 1.0 | 0.0 | 3.0 | 0.0 | 0.0 | 3.0 |
| 07 | 201.0 | 4.0 | 9.0 | 8.0 | 2.0 | 0.0 | 0.0 | 0.0 | 27.0 | 1.0 | 2.0 | 0.0 | 8.0 | 0.0 | 1.0 | 0.0 |
| 08 | 24.0 | 30.0 | 94.0 | 90.0 | 3.0 | 2.0 | 0.0 | 0.0 | 1.0 | 1.0 | 6.0 | 4.0 | 0.0 | 3.0 | 4.0 | 1.0 |
| 09 | 6.0 | 12.0 | 1.0 | 9.0 | 11.0 | 11.0 | 0.0 | 14.0 | 28.0 | 29.0 | 18.0 | 29.0 | 20.0 | 34.0 | 9.0 | 32.0 |
| 10 | 4.0 | 32.0 | 22.0 | 7.0 | 9.0 | 66.0 | 9.0 | 2.0 | 5.0 | 10.0 | 11.0 | 2.0 | 4.0 | 66.0 | 9.0 | 5.0 |
| 11 | 7.0 | 5.0 | 3.0 | 7.0 | 39.0 | 130.0 | 3.0 | 2.0 | 32.0 | 14.0 | 1.0 | 4.0 | 7.0 | 4.0 | 2.0 | 3.0 |
| 12 | 4.0 | 31.0 | 41.0 | 9.0 | 5.0 | 132.0 | 4.0 | 12.0 | 2.0 | 3.0 | 3.0 | 1.0 | 0.0 | 8.0 | 2.0 | 6.0 |
| 13 | 2.0 | 3.0 | 1.0 | 5.0 | 60.0 | 32.0 | 20.0 | 62.0 | 28.0 | 5.0 | 13.0 | 4.0 | 15.0 | 2.0 | 7.0 | 4.0 |
PWM
| AA | AC | AG | AT | CA | CC | CG | CT | GA | GC | GG | GT | TA | TC | TG | TT | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 01 | 1.037 | -1.351 | -0.972 | 0.089 | 0.412 | -1.351 | -0.585 | 0.089 | 1.118 | -1.967 | -2.522 | -0.09 | 0.451 | -2.522 | 0.286 | -0.157 |
| 02 | -1.967 | -1.612 | 2.116 | -1.144 | -3.875 | -3.875 | -0.391 | -3.875 | -3.875 | -2.522 | 0.773 | -2.522 | -2.522 | -2.522 | 1.296 | -1.967 |
| 03 | -3.875 | -3.875 | -1.612 | -3.875 | -2.522 | -3.875 | -1.612 | -2.522 | 1.016 | -2.522 | 2.469 | -1.967 | -1.967 | -3.875 | -0.972 | -3.875 |
| 04 | -1.351 | -3.875 | 0.994 | -3.875 | -3.875 | -3.875 | -2.522 | -3.875 | 0.192 | -3.875 | 2.418 | -1.967 | -3.875 | -3.875 | -1.967 | -2.522 |
| 05 | 0.033 | -3.875 | -0.972 | -2.522 | -3.875 | -3.875 | -3.875 | -3.875 | 2.323 | -1.612 | 1.229 | -1.144 | -1.612 | -3.875 | -3.875 | -3.875 |
| 06 | 2.232 | -1.967 | 0.488 | -0.972 | -2.522 | -3.875 | -1.967 | -3.875 | 1.312 | -3.875 | -2.522 | -3.875 | -1.612 | -3.875 | -3.875 | -1.612 |
| 07 | 2.485 | -1.351 | -0.585 | -0.698 | -1.967 | -3.875 | -3.875 | -3.875 | 0.488 | -2.522 | -1.967 | -3.875 | -0.698 | -3.875 | -2.522 | -3.875 |
| 08 | 0.372 | 0.592 | 1.726 | 1.683 | -1.612 | -1.967 | -3.875 | -3.875 | -2.522 | -2.522 | -0.972 | -1.351 | -3.875 | -1.612 | -1.351 | -2.522 |
| 09 | -0.972 | -0.307 | -2.522 | -0.585 | -0.391 | -0.391 | -3.875 | -0.157 | 0.524 | 0.559 | 0.089 | 0.559 | 0.192 | 0.716 | -0.585 | 0.656 |
| 10 | -1.351 | 0.656 | 0.286 | -0.826 | -0.585 | 1.374 | -0.585 | -1.967 | -1.144 | -0.484 | -0.391 | -1.967 | -1.351 | 1.374 | -0.585 | -1.144 |
| 11 | -0.826 | -1.144 | -1.612 | -0.826 | 0.852 | 2.05 | -1.612 | -1.967 | 0.656 | -0.157 | -2.522 | -1.351 | -0.826 | -1.351 | -1.967 | -1.612 |
| 12 | -1.351 | 0.625 | 0.901 | -0.585 | -1.144 | 2.065 | -1.351 | -0.307 | -1.967 | -1.612 | -1.612 | -2.522 | -3.875 | -0.698 | -1.967 | -0.972 |
| 13 | -1.967 | -1.612 | -2.522 | -1.144 | 1.28 | 0.656 | 0.192 | 1.312 | 0.524 | -1.144 | -0.229 | -1.351 | -0.09 | -1.967 | -0.826 | -1.351 |