PWMs for transcription factors
| Model | LOGO | Transcription factor | Model length | Quality | Model rank | Consensus | Model release | Data source | Best auROC (human) | Best auROC (mouse) | Peak sets in benchmark (human) | Peak sets in benchmark (mouse) | Aligned words | TF family | TF subfamily | HGNC | MGI | EntrezGene | UniProt ID | UniProt AC |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NKX21_HUMAN.H11MO.0.A |  |
HUMAN:NKX2-1 | 10 | A | 0 | bWKGAGWGbh | HOCOMOCOv11 | ChIP-Seq | 0.92 | 0.917 | 14 | 9 | 501 | NK-related factors{3.1.2} | NK-2.1{3.1.2.14} | 11825 | 7080 | NKX21_HUMAN | P43699 | |
| NKX21_HUMAN.H11DI.0.A |  |
HUMAN:NKX2-1 | 13 | A | 0 | nbYKGAGWGbnhn | HOCOMOCOv11 | ChIP-Seq | 0.938 | 0.927 | 14 | 9 | 501 | NK-related factors{3.1.2} | NK-2.1{3.1.2.14} | 11825 | 7080 | NKX21_HUMAN | P43699 | |
| NKX21_MOUSE.H11MO.0.A |  |
MOUSE:Nkx2-1 | 10 | A | 0 | bbWKGAGWGb | HOCOMOCOv11 | ChIP-Seq | 0.923 | 0.92 | 14 | 9 | 500 | NK-related factors{3.1.2} | NK-2.1{3.1.2.14} | 108067 | 21869 | NKX21_MOUSE | P50220 | |
| NKX21_MOUSE.H11DI.0.A | 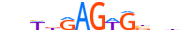 |
MOUSE:Nkx2-1 | 13 | A | 0 | nbWbRAGWGbbhn | HOCOMOCOv11 | ChIP-Seq | 0.905 | 0.937 | 14 | 9 | 502 | NK-related factors{3.1.2} | NK-2.1{3.1.2.14} | 108067 | 21869 | NKX21_MOUSE | P50220 |