PWMs for transcription factors
| Model | LOGO | Transcription factor | Model length | Quality | Model rank | Consensus | Model release | Data source | Best auROC (human) | Best auROC (mouse) | Peak sets in benchmark (human) | Peak sets in benchmark (mouse) | Aligned words | TF family | TF subfamily | HGNC | MGI | EntrezGene | UniProt ID | UniProt AC |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| MGAP_HUMAN.H11MO.0.D |  |
HUMAN:MGA | 16 | D | 0 | RRGTGTKAnWTWTMRC | HOCOMOCOv10 | HT-SELEX | 6178 | bHLH-ZIP factors{1.2.6}; TBX6-related factors{6.5.5} |
Mad-like factors{1.2.6.7}; MGA (MAD5){6.5.5.0.2} |
14010 | 23269 | MGAP_HUMAN | Q8IWI9 | |||||
| MXI1_HUMAN.H11MO.0.A | 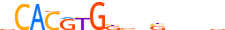 |
HUMAN:MXI1 | 15 | A | 0 | vCACRYGKvnSbbbv | HOCOMOCOv10 | ChIP-Seq | 0.878 | 0.973 | 10 | 5 | 500 | bHLH-ZIP factors{1.2.6} | Mad-like factors{1.2.6.7} | 7534 | 4601 | MXI1_HUMAN | P50539 | |
| MXI1_HUMAN.H11MO.1.A |  |
HUMAN:MXI1 | 11 | A | 1 | SMCACRTGbhb | HOCOMOCOv11 | ChIP-Seq | 0.871 | 0.96 | 10 | 5 | 500 | bHLH-ZIP factors{1.2.6} | Mad-like factors{1.2.6.7} | 7534 | 4601 | MXI1_HUMAN | P50539 | |
| MXI1_HUMAN.H11DI.0.A |  |
HUMAN:MXI1 | 23 | A | 0 | nvvvCAYGTGSvnvvnbvvvnbn | HOCOMOCOv10 | ChIP-Seq | 0.897 | 0.981 | 10 | 5 | 500 | bHLH-ZIP factors{1.2.6} | Mad-like factors{1.2.6.7} | 7534 | 4601 | MXI1_HUMAN | P50539 | |
| MXI1_MOUSE.H11MO.0.A | 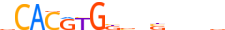 |
MOUSE:Mxi1 | 15 | A | 0 | vCACRYGKvnSbbbv | HOCOMOCOv10 | ChIP-Seq | 0.878 | 0.973 | 10 | 5 | 500 | bHLH-ZIP factors{1.2.6} | Mad-like factors{1.2.6.7} | 97245 | 17859 | MXI1_MOUSE | P50540 | |
| MXI1_MOUSE.H11MO.1.A |  |
MOUSE:Mxi1 | 8 | A | 1 | hCRCGTGS | HOCOMOCOv11 | ChIP-Seq | 0.856 | 0.973 | 10 | 5 | 510 | bHLH-ZIP factors{1.2.6} | Mad-like factors{1.2.6.7} | 97245 | 17859 | MXI1_MOUSE | P50540 | |
| MXI1_MOUSE.H11DI.0.A |  |
MOUSE:Mxi1 | 23 | A | 0 | nvvvCAYGTGSvnvvnbvvvnbn | HOCOMOCOv10 | ChIP-Seq | 0.897 | 0.981 | 10 | 5 | 500 | bHLH-ZIP factors{1.2.6} | Mad-like factors{1.2.6.7} | 97245 | 17859 | MXI1_MOUSE | P50540 |