PWMs for transcription factors
| Model | LOGO | Transcription factor | Model length | Quality | Model rank | Consensus | Model release | Data source | Best auROC (human) | Best auROC (mouse) | Peak sets in benchmark (human) | Peak sets in benchmark (mouse) | Aligned words | TF family | TF subfamily | HGNC | MGI | EntrezGene | UniProt ID | UniProt AC |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| LHX2_HUMAN.H11MO.0.A |  |
HUMAN:LHX2 | 12 | A | 0 | bbYTAATTRvhb | HOCOMOCOv11 | ChIP-Seq | 0.826 | 0.931 | 5 | 12 | 430 | HD-LIM factors{3.1.5} | Lhx-2-like factors{3.1.5.3} | 6594 | 9355 | LHX2_HUMAN | P50458 | |
| LHX9_HUMAN.H11MO.0.D | 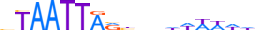 |
HUMAN:LHX9 | 18 | D | 0 | bTAATTARbnhhWWWWdd | HOCOMOCOv10 | HT-SELEX | 4409 | HD-LIM factors{3.1.5} | Lhx-2-like factors{3.1.5.3} | 14222 | 56956 | LHX9_HUMAN | Q9NQ69 | |||||
| LHX2_HUMAN.H11DI.0.A |  |
HUMAN:LHX2 | 12 | A | 0 | nYTAATKRbbhn | HOCOMOCOv11 | ChIP-Seq | 0.831 | 0.938 | 5 | 12 | 501 | HD-LIM factors{3.1.5} | Lhx-2-like factors{3.1.5.3} | 6594 | 9355 | LHX2_HUMAN | P50458 | |
| LHX2_MOUSE.H11MO.0.A |  |
MOUSE:Lhx2 | 10 | A | 0 | nhTAATKAGn | HOCOMOCOv11 | ChIP-Seq | 0.814 | 0.938 | 5 | 12 | 501 | HD-LIM factors{3.1.5} | Lhx-2-like factors{3.1.5.3} | 96785 | 16870 | LHX2_MOUSE | Q9Z0S2 | |
| LHX2_MOUSE.H11DI.0.A |  |
MOUSE:Lhx2 | 11 | A | 0 | nhTAATKAGvn | HOCOMOCOv11 | ChIP-Seq | 0.804 | 0.934 | 5 | 12 | 531 | HD-LIM factors{3.1.5} | Lhx-2-like factors{3.1.5.3} | 96785 | 16870 | LHX2_MOUSE | Q9Z0S2 |