PWMs for transcription factors
| Model | LOGO | Transcription factor | Model length | Quality | Model rank | Consensus | Model release | Data source | Best auROC (human) | Best auROC (mouse) | Peak sets in benchmark (human) | Peak sets in benchmark (mouse) | Aligned words | TF family | TF subfamily | HGNC | MGI | EntrezGene | UniProt ID | UniProt AC |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NFAC1_HUMAN.H11MO.0.B |  |
HUMAN:NFATC1 | 15 | B | 0 | vWGGAAAvdvWRvvd | HOCOMOCOv11 | ChIP-Seq | 0.868 | 0.795 | 7 | 8 | 500 | NFAT-related factors{6.1.3} | NFATc1{6.1.3.0.1} | 7775 | 4772 | NFAC1_HUMAN | O95644 | |
| NFAC1_HUMAN.H11MO.1.B |  |
HUMAN:NFATC1 | 10 | B | 1 | vvWGGAAARn | HOCOMOCOv11 | ChIP-Seq | 0.792 | 0.828 | 7 | 8 | 500 | NFAT-related factors{6.1.3} | NFATc1{6.1.3.0.1} | 7775 | 4772 | NFAC1_HUMAN | O95644 | |
| NFAC1_HUMAN.H11DI.0.A | 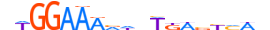 |
HUMAN:NFATC1 | 18 | A | 0 | dWGGAARndnTKWvYMWn | HOCOMOCOv11 | ChIP-Seq | 0.812 | 0.954 | 7 | 8 | 500 | NFAT-related factors{6.1.3} | NFATc1{6.1.3.0.1} | 7775 | 4772 | NFAC1_HUMAN | O95644 | |
| NFAC1_MOUSE.H11MO.0.A |  |
MOUSE:Nfatc1 | 15 | A | 0 | WGGAAAndnTKWvhM | HOCOMOCOv11 | ChIP-Seq | 0.714 | 0.955 | 7 | 8 | 445 | NFAT-related factors{6.1.3} | NFATc1{6.1.3.0.1} | 102469 | NFAC1_MOUSE | O88942 | ||
| NFAC1_MOUSE.H11MO.1.A |  |
MOUSE:Nfatc1 | 11 | A | 1 | vvvWGGAAAvn | HOCOMOCOv11 | ChIP-Seq | 0.689 | 0.908 | 7 | 8 | 500 | NFAT-related factors{6.1.3} | NFATc1{6.1.3.0.1} | 102469 | NFAC1_MOUSE | O88942 | ||
| NFAC1_MOUSE.H11DI.0.A |  |
MOUSE:Nfatc1 | 18 | A | 0 | vWGGAARvdnWKWvWSRn | HOCOMOCOv11 | ChIP-Seq | 0.726 | 0.963 | 7 | 8 | 508 | NFAT-related factors{6.1.3} | NFATc1{6.1.3.0.1} | 102469 | NFAC1_MOUSE | O88942 |