PWMs for transcription factors
| Model | LOGO | Transcription factor | Model length | Quality | Model rank | Consensus | Model release | Data source | Best auROC (human) | Best auROC (mouse) | Peak sets in benchmark (human) | Peak sets in benchmark (mouse) | Aligned words | TF family | TF subfamily | HGNC | MGI | EntrezGene | UniProt ID | UniProt AC |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| STA5A_HUMAN.H11MO.0.A |  |
HUMAN:STAT5A | 10 | A | 0 | TTCYARGAAd | HOCOMOCOv11 | ChIP-Seq | 0.932 | 0.992 | 24 | 127 | 500 | STAT factors{6.2.1} | STAT5A{6.2.1.0.5} | 11366 | 6776 | STA5A_HUMAN | P42229 | |
| STA5B_HUMAN.H11MO.0.A |  |
HUMAN:STAT5B | 12 | A | 0 | nTTCYTRGAAdh | HOCOMOCOv11 | ChIP-Seq | 0.965 | 0.957 | 16 | 31 | 501 | STAT factors{6.2.1} | STAT5B{6.2.1.0.6} | 11367 | 6777 | STA5B_HUMAN | P51692 | |
| STAT1_HUMAN.H11MO.0.A |  |
HUMAN:STAT1 | 19 | A | 0 | hbhnYTTCCnRGAARhShb | HOCOMOCOv11 | ChIP-Seq | 0.987 | 0.91 | 37 | 128 | 500 | STAT factors{6.2.1} | STAT1{6.2.1.0.1} | 11362 | 6772 | STAT1_HUMAN | P42224 | |
| STAT1_HUMAN.H11MO.1.A |  |
HUMAN:STAT1 | 19 | A | 1 | vRRRAAhnGAAAShdRRdn | HOCOMOCOv11 | ChIP-Seq | 0.992 | 0.962 | 37 | 128 | 500 | STAT factors{6.2.1} | STAT1{6.2.1.0.1} | 11362 | 6772 | STAT1_HUMAN | P42224 | |
| STAT2_HUMAN.H11MO.0.A |  |
HUMAN:STAT2 | 19 | A | 0 | dvRRAAhhGAAASYdRRRv | HOCOMOCOv11 | ChIP-Seq | 0.98 | 0.994 | 11 | 8 | 500 | STAT factors{6.2.1} | STAT2{6.2.1.0.2} | 11363 | 6773 | STAT2_HUMAN | P52630 | |
| STAT3_HUMAN.H11MO.0.A |  |
HUMAN:STAT3 | 12 | A | 0 | bvhTTCCCRKAA | HOCOMOCOv11 | ChIP-Seq | 0.953 | 0.949 | 36 | 82 | 500 | STAT factors{6.2.1} | STAT3{6.2.1.0.3} | 11364 | 6774 | STAT3_HUMAN | P40763 | |
| STAT4_HUMAN.H11MO.0.A | 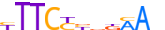 |
HUMAN:STAT4 | 11 | A | 0 | YTTCYYdKMAv | HOCOMOCOv11 | ChIP-Seq | 0.935 | 0.887 | 7 | 10 | 500 | STAT factors{6.2.1} | STAT4{6.2.1.0.4} | 11365 | 6775 | STAT4_HUMAN | Q14765 | |
| STAT6_HUMAN.H11MO.0.B |  |
HUMAN:STAT6 | 11 | B | 0 | TTCYbWRGAAv | HOCOMOCOv11 | ChIP-Seq | 0.766 | 0.912 | 19 | 24 | 502 | STAT factors{6.2.1} | STAT6{6.2.1.0.7} | 11368 | 6778 | STAT6_HUMAN | P42226 | |
| STA5A_HUMAN.H11DI.0.A |  |
HUMAN:STAT5A | 25 | A | 0 | nddnndWKMhdRGAAddRvvnvddn | HOCOMOCOv11 | ChIP-Seq | 0.949 | 0.984 | 24 | 127 | 503 | STAT factors{6.2.1} | STAT5A{6.2.1.0.5} | 11366 | 6776 | STA5A_HUMAN | P42229 | |
| STA5B_HUMAN.H11DI.0.A |  |
HUMAN:STAT5B | 12 | A | 0 | dhTTCYTRGAAn | HOCOMOCOv11 | ChIP-Seq | 0.963 | 0.963 | 16 | 31 | 508 | STAT factors{6.2.1} | STAT5B{6.2.1.0.6} | 11367 | 6777 | STA5B_HUMAN | P51692 | |
| STAT1_HUMAN.H11DI.0.A |  |
HUMAN:STAT1 | 25 | A | 0 | bbvbTTCCYRGAARhvnnnbbvnnn | HOCOMOCOv11 | ChIP-Seq | 0.989 | 0.917 | 37 | 128 | 500 | STAT factors{6.2.1} | STAT1{6.2.1.0.1} | 11362 | 6772 | STAT1_HUMAN | P42224 | |
| STAT2_HUMAN.H11DI.0.A |  |
HUMAN:STAT2 | 20 | A | 0 | ndvRRAAhnGAAASYdRRWn | HOCOMOCOv11 | ChIP-Seq | 0.982 | 0.995 | 11 | 8 | 500 | STAT factors{6.2.1} | STAT2{6.2.1.0.2} | 11363 | 6773 | STAT2_HUMAN | P52630 | |
| STAT3_HUMAN.H11DI.0.A | 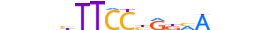 |
HUMAN:STAT3 | 18 | A | 0 | nbbvbTTCChRKMAvnbn | HOCOMOCOv10 | ChIP-Seq | 0.956 | 0.952 | 36 | 82 | 436 | STAT factors{6.2.1} | STAT3{6.2.1.0.3} | 11364 | 6774 | STAT3_HUMAN | P40763 | |
| STAT4_HUMAN.H11DI.0.A |  |
HUMAN:STAT4 | 22 | A | 0 | nvnnnbbnbTTCCYdKMAdnnn | HOCOMOCOv10 | ChIP-Seq | 0.958 | 0.942 | 7 | 10 | 502 | STAT factors{6.2.1} | STAT4{6.2.1.0.4} | 11365 | 6775 | STAT4_HUMAN | Q14765 | |
| STAT6_HUMAN.H11DI.0.A |  |
HUMAN:STAT6 | 15 | A | 0 | ndvRvAGGAARbvdn | HOCOMOCOv11 | ChIP-Seq | 0.874 | 0.834 | 19 | 24 | 506 | STAT factors{6.2.1} | STAT6{6.2.1.0.7} | 11368 | 6778 | STAT6_HUMAN | P42226 | |
| STA5A_MOUSE.H11MO.0.A |  |
MOUSE:Stat5a | 10 | A | 0 | TTCYARGAAv | HOCOMOCOv11 | ChIP-Seq | 0.928 | 0.992 | 24 | 127 | 500 | STAT factors{6.2.1} | STAT5A{6.2.1.0.5} | 103036 | 20850 | STA5A_MOUSE | P42230 | |
| STA5B_MOUSE.H11MO.0.A |  |
MOUSE:Stat5b | 12 | A | 0 | nTTCYTRGAAdh | HOCOMOCOv11 | ChIP-Seq | 0.965 | 0.957 | 16 | 31 | 501 | STAT factors{6.2.1} | STAT5B{6.2.1.0.6} | 103035 | 20851 | STA5B_MOUSE | P42232 | |
| STAT1_MOUSE.H11MO.0.A |  |
MOUSE:Stat1 | 22 | A | 0 | dRRRAAhdGAAAvhvMvnvhdd | HOCOMOCOv11 | ChIP-Seq | 0.989 | 0.958 | 37 | 128 | 500 | STAT factors{6.2.1} | STAT1{6.2.1.0.1} | 103063 | STAT1_MOUSE | P42225 | ||
| STAT1_MOUSE.H11MO.1.A |  |
MOUSE:Stat1 | 21 | A | 1 | bvdhTKMhRGGAARhvRRvhv | HOCOMOCOv11 | ChIP-Seq | 0.976 | 0.951 | 37 | 128 | 500 | STAT factors{6.2.1} | STAT1{6.2.1.0.1} | 103063 | STAT1_MOUSE | P42225 | ||
| STAT2_MOUSE.H11MO.0.A |  |
MOUSE:Stat2 | 19 | A | 0 | dvRRAAhhGAAASYdRRRv | HOCOMOCOv11 | ChIP-Seq | 0.98 | 0.994 | 11 | 8 | 500 | STAT factors{6.2.1} | STAT2{6.2.1.0.2} | 103039 | STAT2_MOUSE | Q9WVL2 | ||
| STAT3_MOUSE.H11MO.0.A |  |
MOUSE:Stat3 | 12 | A | 0 | bdYTTCCMRKMA | HOCOMOCOv11 | ChIP-Seq | 0.94 | 0.941 | 36 | 82 | 500 | STAT factors{6.2.1} | STAT3{6.2.1.0.3} | 103038 | 20848 | STAT3_MOUSE | P42227 | |
| STAT4_MOUSE.H11MO.0.A |  |
MOUSE:Stat4 | 10 | A | 0 | YTTCYhdKMM | HOCOMOCOv11 | ChIP-Seq | 0.932 | 0.914 | 7 | 10 | 475 | STAT factors{6.2.1} | STAT4{6.2.1.0.4} | 103062 | 20849 | STAT4_MOUSE | P42228 | |
| STAT6_MOUSE.H11MO.0.A |  |
MOUSE:Stat6 | 14 | A | 0 | bTTCYhvRGAAvhv | HOCOMOCOv11 | ChIP-Seq | 0.751 | 0.932 | 19 | 24 | 407 | STAT factors{6.2.1} | STAT6{6.2.1.0.7} | 103034 | 20852 | STAT6_MOUSE | P52633 | |
| STA5A_MOUSE.H11DI.0.A |  |
MOUSE:Stat5a | 13 | A | 0 | ndnTTCYMRGAAn | HOCOMOCOv11 | ChIP-Seq | 0.927 | 0.993 | 24 | 127 | 500 | STAT factors{6.2.1} | STAT5A{6.2.1.0.5} | 103036 | 20850 | STA5A_MOUSE | P42230 | |
| STA5B_MOUSE.H11DI.0.A |  |
MOUSE:Stat5b | 13 | A | 0 | nnTTCYTRGAAvn | HOCOMOCOv11 | ChIP-Seq | 0.966 | 0.965 | 16 | 31 | 501 | STAT factors{6.2.1} | STAT5B{6.2.1.0.6} | 103035 | 20851 | STA5B_MOUSE | P42232 | |
| STAT1_MOUSE.H11DI.0.A |  |
MOUSE:Stat1 | 24 | A | 0 | ndRRRAAhhGAAASWvRWdvnvdn | HOCOMOCOv11 | ChIP-Seq | 0.988 | 0.966 | 37 | 128 | 506 | STAT factors{6.2.1} | STAT1{6.2.1.0.1} | 103063 | STAT1_MOUSE | P42225 | ||
| STAT2_MOUSE.H11DI.0.A |  |
MOUSE:Stat2 | 21 | A | 0 | nvvRRAAhnGAAASYRRRdnn | HOCOMOCOv11 | ChIP-Seq | 0.981 | 0.994 | 11 | 8 | 509 | STAT factors{6.2.1} | STAT2{6.2.1.0.2} | 103039 | STAT2_MOUSE | Q9WVL2 | ||
| STAT3_MOUSE.H11DI.0.A |  |
MOUSE:Stat3 | 20 | A | 0 | nhbvbTTMCYRKvAnbnnbn | HOCOMOCOv11 | ChIP-Seq | 0.95 | 0.946 | 36 | 82 | 509 | STAT factors{6.2.1} | STAT3{6.2.1.0.3} | 103038 | 20848 | STAT3_MOUSE | P42227 | |
| STAT4_MOUSE.H11DI.0.A |  |
MOUSE:Stat4 | 22 | A | 0 | nvnnnbbnbTTCCYdKMAdnnn | HOCOMOCOv10 | ChIP-Seq | 0.958 | 0.942 | 7 | 10 | 502 | STAT factors{6.2.1} | STAT4{6.2.1.0.4} | 103062 | 20849 | STAT4_MOUSE | P42228 | |
| STAT6_MOUSE.H11DI.0.A |  |
MOUSE:Stat6 | 18 | A | 0 | nbdnTTCYWvRGAAnnnn | HOCOMOCOv11 | ChIP-Seq | 0.771 | 0.951 | 19 | 24 | 492 | STAT factors{6.2.1} | STAT6{6.2.1.0.7} | 103034 | 20852 | STAT6_MOUSE | P52633 |